Mass Spectrometry Wields its Might in the Battle Against COVID-19
Mass Spec has advanced to embrace some of the most challenging applications in life sciences research.
An evolution of technology has brought everything from single-cell proteomics to systems-level metabolomics into reality.
Faced with the ongoing challenges of COVID-19, mass spec is once again wielding its might in the battle against the disease. Here is a brief look at some of the method development and collaborative efforts that are part of the charge.
MS-Based Virus Detection in Saliva
Diagnostics testing for SARS-CoV-2 virus has been challenged in multiple ways, and new techniques are adding vital ammunition to the growing cache of testing technologies.
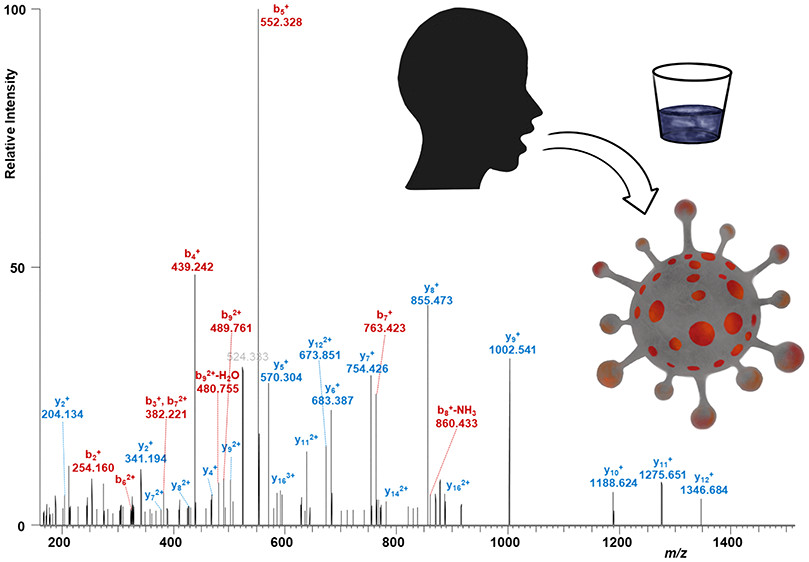
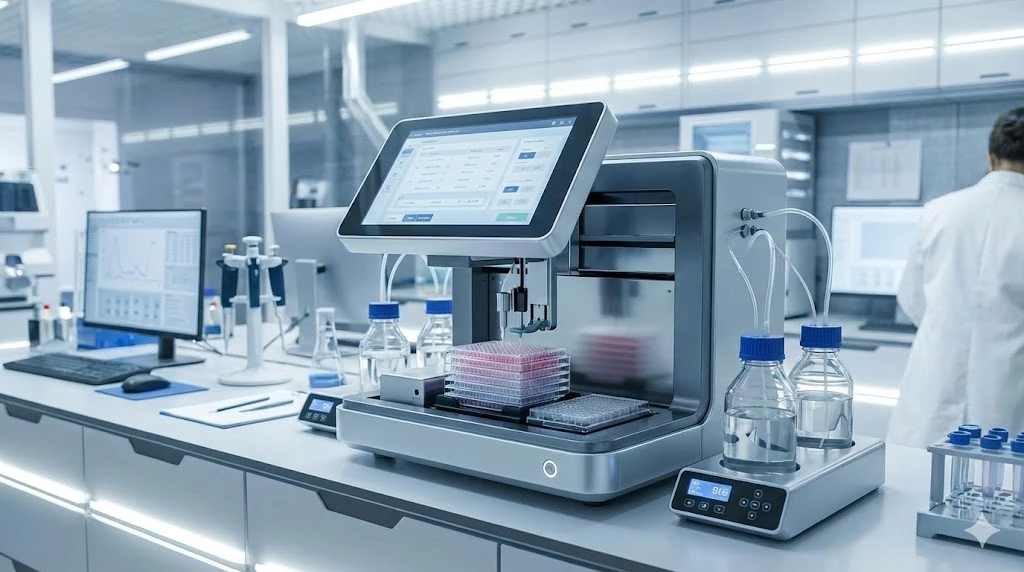
Researchers recently developed a MS-based test that uses gargle samples from patients, rather than throat of nasal samples. As opposed to detecting RNA genetic material, the MS approach detects virus-specific proteins present in easy-to-obtain patient saliva.
The protocol is relatively straightforward, involving acetone precipitation and tryptic digestion of proteins, followed by MS analysis. LC-MS/MS is performed using a nano-HPLC system coupled with an Orbitrap Fusion Hybrid mass spectrometer. Samples are first loaded on a trapping column, washed, and then subjected to a separation column prior to subjecting to nano-ESI. The method proof of concept detected unique peptides derived from the SARS-CoV-2 protein in confirmed COVID-19 patient samples.
The pros of the approach include definitive detection of SARS-CoV0-2 nucleocapsid protein among a high background of human proteins and peptides. Saliva is much more easily obtainable than throat or nasal swabs, and relative viral load can be deduced using this method adding valuable sensitivity potential. Cons include the fact that sample preparation involves multiple steps including glycoprotein removal, thereby limiting the high-throughput potential. Long analysis times and the need for more extensive protein identification, beyond the single nucleocapsid peptide in the study, are future areas for improvement as well.
MS-Based Validation of Viral Marker Peptides
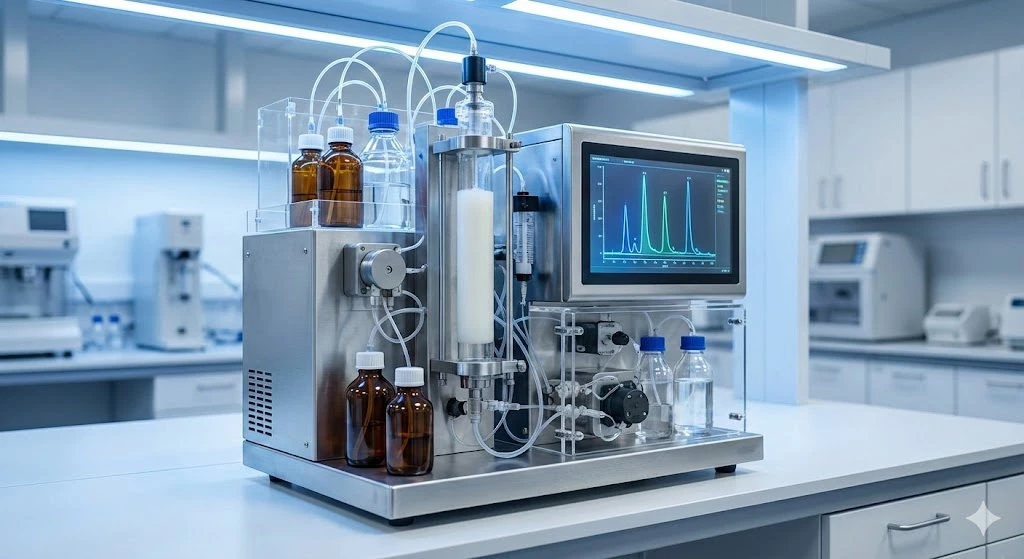
Another proof of concept research study took aim at identifying the most specific and promising marker peptides to include in a panel for definitive SARS-CoV-2 detection.
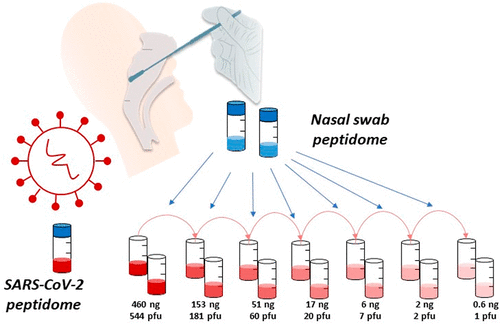
The work used nasal swabs spiked with differing quantities of purified viral material to validate the novel nano-LC-MS/MS method. This technique was then successfully used on COVID-19 clinical samples.
Results identified two unique peptides from the SARS-CoV-2 nucleocapsid protein as highly specific and sensitive viral markers. Additionally, results showed the peptide signals can be obtained within a 3-minute window in the tested conditions, providing a time-efficient multi-peptide diagnostic approach ideal for further clinical development.
Mass Spectrometry Coalitions Against COVID-19
These studies are examples of powerful LC-MS technologies applied to public health emergencies such as the ongoing COVID-19 pandemic.
Large-scale collaborations have brought experts together from diverse fields to channel mass spec technical expertise towards the COVID-19 challenge. As part of their mission, the COVID-19 Mass Spectrometry coalition aims “to use mass spec to reduce the harm caused by the SARS-Cov-2 virus” by sharing “sample collection and processing protocols, and all data that is obtained to maximize benefit for all”.
The power of mass spec is being applied to projects including:
- Mapping the viral antigens in blood and other biofluids to inform serological testing
- Assaying and performing quality control on compounds produced for serological tests
- Applying MS to inform vaccine and therapeutic development efforts
- Developing methods to determine disease progression and viral transmission in the environment
Mass Spectrometry Industry Partners Join the Fight
Those in the mass spec industry have assembled resources to arm those in the fight as well.
Bruker announced support for a COVID-19-NMR Consortium meant to investigate the structure, dynamics, function, and binding of key SARS-CoZ-2 proteins using nuclear magnetic resonance (NMR). The group’s website currently indicates 170 collaborators from 15 different countries, coming together on 17 RNA and 12 protein projects. With expertise in many different areas of analytical technology, Bruker is no doubt leveraging its considerable talent pool to provide the means for this endeavor.
Waters launched a COVID-19 Resource Hub with the purpose of enhancing human health in the face of the COVID-19 challenge. Expert teams have pulled together resources to provide up to minute news, applications, and on-demand support or researchers. Among this information is insight into the future of science and the increasing role that mass spec will serve in diagnostics and molecular medicine.
As discussed previously, other companies including MOBILion and Ionicon have announced new methods and collaborations to characterize the SARS-CoV-2 spike glycoprotein and breath-based viral detection strategies, respectively, both integrating mass spectrometry as the main analytical platform. A host of additional companies have assembled their technologies as well.
Outlook
As a science community, we have never been more prepared to face such a challenge as COVID-19. The unique aspects of this viral disease have required mobilization of advanced technologies to the frontlines of research and diagnostic development. Innovative mass spec approaches will ultimately prove essential in fighting back this pandemic and for confronting emerging infectious diseases in the future.
View Mass Spectrometer Listings on LabX.com
View HPLC Systems Listings on LabX.com
Image 1 Credit:
Mass Spectrometric Identification of SARS-CoV-2 Proteins from Gargle Solution Samples of COVID-19 Patients
Christian Ihling, Dirk Tänzler, Sven Hagemann, Astrid Kehlen, Stefan Hüttelmaier, Christian Arlt, and Andrea Sinz
Journal of Proteome Research Article ASAP
DOI: 10.1021/acs.jproteome.0c00280
Image 2 Credit:
Proteotyping SARS-CoV-2 Virus from Nasopharyngeal Swabs: A Proof-of-Concept Focused on a 3 Min Mass Spectrometry Window
Duarte Gouveia, Guylaine Miotello, Fabrice Gallais, Jean-Charles Gaillard, Stéphanie Debroas, Laurent Bellanger, Jean-Philippe Lavigne, Albert Sotto, Lucia Grenga, Olivier Pible, and Jean Armengaud
Journal of Proteome Research Article ASAP
DOI: 10.1021/acs.jproteome.0c00535