Progress Along the Frontier of Next-Generation Proteomics
Next-Generation Genomics
The development of high-performance sequencing technologies provoked a paradigm shift in DNA and nucleic acid sequencing analysis and the birth of Next-Generation Sequencing (NGS). This is turn, motivated the large-scale sequencing and annotation of whole-genomes, and the advent of the molecular genomics and genome engineering fields. As a result of these monumental achievements, progress along multiple fronts led a new age of molecular medicine – with NGS in vitro diagnostics and precision medicine at the forefront.
Next-Generation Proteomics
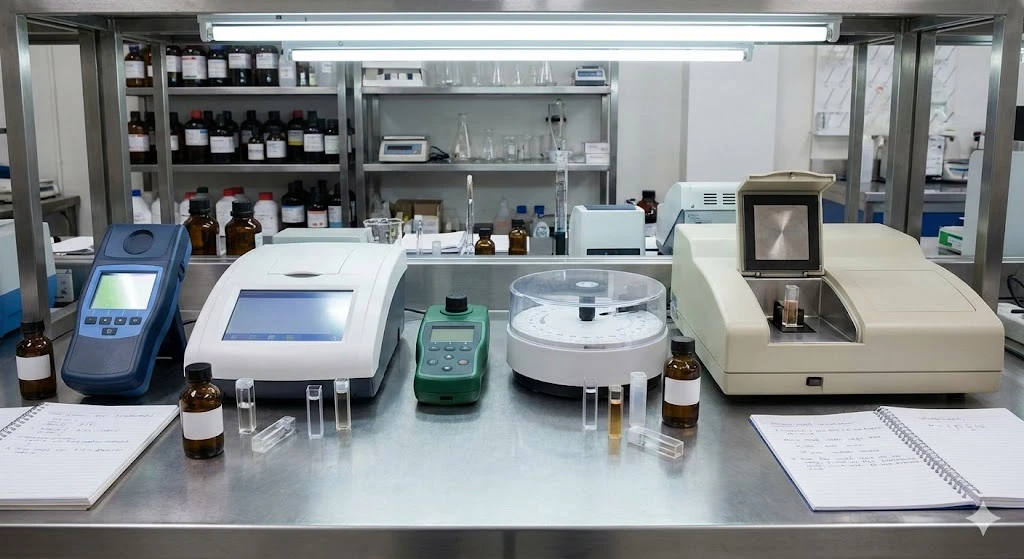
Biological mass spectrometry is witnessing a similar paradigm shift. Technological advancements in speed, sensitivity, and resolution have enabled the measurement of minute differences in protein characteristics in large sample groups and in complex backgrounds. Enhancements in sample preparation methods and the capture of large data streams have complemented this new era in proteomics.
The concept of Next-Generation Proteomics aims to dissect distinct protein profiles in small sample sizes and in native tissues – including those at the single-cell level. These abilities will enable the detection of minute differences underlying cancerous cells versus benign, infected cells versus healthy, and other important disease distinctions. As the concept matures, next-gen proteomics will produce a new layer of insight to complement NGS in the realization of true precision medicine.
Several advanced technologies showcased at the American Society for Mass Spectrometry annual meeting are indicative of progress along the frontier of next-generation proteomics.
Innovations in data capture and annotation
With the ability to now create vastly abundant peptide species from often complex and large sample sets, the limiting factor has become comprehensive data capture and meaningful analysis. SCIEX originally pioneered the SWATH technology as a novel data-independent acquisition (DIA) strategy to complement multi-reaction monitoring (MRM) and data-dependent acquisition (DDA) techniques.
How does SWATH work?
SWATH excels by the use of an expanded mass isolation window (Q1) and the stepwise movement of this window across a wide mass range, covering a comprehensive distribution of peptides and full-scan MS-MS data. Post-acquisition, ion chromatograms for each fragment ion are processed and quantification is performed. A digital record of all peptide species and their relevant abundance is recorded and stored in a spectral ion library – essentially a comprehensive reference for ongoing experiments on the proteome of interest.
What is Scanning SWATH and what can it do?
The new Scanning SWATH acquisition technology, introduced at ASMS this year and built into the new TripleTOF 6600+ LC-MS/MS system, captures more detail about proteins and potential markers, effectively creating a detailed digital fingerprint of the proteome.
How is Scanning SWATH different?
The technology uses a sliding Q1 mass selection window, increased mass range coverage and depth, and high-confidence fragment/peptide matching. The speed and resolution of the previous SWATH “stepping” is increased to essentially enable scanning of complete peptide profiles and high-resolution proteome analysis. Coupled with the now cloud-based OneOmics software, big data generated from these experiments can be integrated into large metabolomics and genomics studies – and actionable results can be obtained.
Innovations in imaging proteomics
The capture of as much information as possible to complete the spatial and molecular picture of cells and tissues is a goal of next-gen proteomics imaging. Bruker introduced the new timsTOF flex mass spectrometer at ASMS this year that promises to further this objective.
The timsTOF flex has a software switchable MALDI source, which together with the traditional ESI source (derived from the previous timsTOF Pro) enables “spatially-resolved omics, or SpatialOMx”, on a single instrument. The 10kHz SmartBeam 3D laser provides high-fidelity, rapid, label-free MALDI imaging with high-spatial resolution. Importantly, ESI analysis is preserved, adding molecular level characterization as another dimension – the concept of 4D imaging proteomics.
The platform is geared to allow specific targeting of cell sub-populations for subsequent ESI-TIMS/PASEF-based DDA or DIA analysis. Applications include lipidomics and metabolomics investigations of discrete cell states, such as cancer, autoimmunity, infectious disease and other disease scenarios where cell-level resolution is advantageous. Distinguishing disease from disease-free tissues - at the single-cell level - is a goal inline with next-generation proteomics.
The outlook for next-generation proteomics
Advancements in high-content data capture, such as the SWATH and TIMS/PASEF technologies, have added depth to proteomics analysis, complementing advancements in speed, high-accuracy, high-resolution. Acquiring the most accurate, comprehensive, and definitive information from the most concise sample possible is the vision for the future. Many exciting developments this year and the next may bring this closer to reality.