4 Features of PCR that Underlie Its Power and Potential
From humble roots to noble humanitarian cause, PCR has left an indelible impact on the world of science
Over 35 years ago, the polymerase chain reaction (PCR) technique was born. The brainchild of the late Kary Mullins, PCR went on to change the landscape of Molecular Biology and win Mullins the Nobel Prize.
In the modern day, PCR and it’s influence on molecular biology, medicine, and humanity for that matter cannot be overstated. Through somewhat humble beginnings, PCR has risen to become arguably the most important technology in combatting challenges such as the SARS-CoV-2 pandemic.
What features of PCR make it so well suited for the extensive range of applications, and what does the future hold for PCR moving forward? Here we look at four features that underlie the power and the potential of the technique.
Sensitivity and Specificity
For review, specific and minute amounts of DNA can be identified and the underlying genetic code can be revealed using PCR. In the process, single-strand primers locate and bind to specific sequences on sample DNA which has been denatured into the two complementary strands. Subsequent annealing, extension, and melting steps amplify the target region, enabling detection using probes. More in-depth description of the technique is offered here.
The specificity of PCR can be controlled by the chemical nature of the primer set, the performance of the PCR polymerase, and by control of temperature and duration of the cycling steps.
Primer Length and Sequence
PCR primers can be produced to bind tightly to the sequence of a single strand of the target DNA. Increasing the primer length can increase this affinity. Whether short sequence elements will need to be added to the PCR product, such as introduction of a restriction enzyme cleavage site, can also influence primer length and affinity.
PCR Polymerase Activity
The performance of the reaction mix and the DNA polymerase directly correlates to the fidelity in which the DNA sequence is amplified. High fidelity polymerase may be ideal for challenging sequences. Standard polymerase may be more suitable for workhorse PCR for rapid detection or diagnostics. Error-prone PCR can be useful in creating genetic mutations for research purposes. For each application there is a suitable polymerase that plays a direct role in the specificity of the PCR.
Thermocycling Conditions
 All of these factors are considered in choosing the right thermocycling conditions. Many computational tools are available to help guide the design of suitable PCR conditions. Using such tools, input of the target DNA and primer sequences enables calculation of ideal annealing, extension, and denaturing temperatures. Features such as the length of target sequence will affect the duration of each cycle.
All of these factors are considered in choosing the right thermocycling conditions. Many computational tools are available to help guide the design of suitable PCR conditions. Using such tools, input of the target DNA and primer sequences enables calculation of ideal annealing, extension, and denaturing temperatures. Features such as the length of target sequence will affect the duration of each cycle.
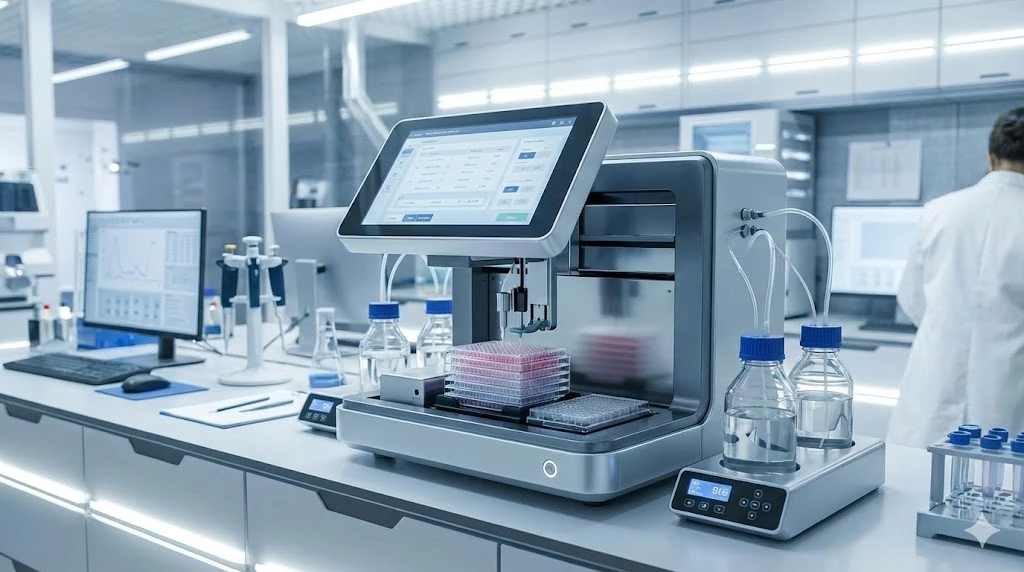
Although there are many further details involved, the point to be made is PCR has proven highly amenable to modification by tuning the specificity and sensitivity according to the requirements of the user. These attributes alone have made the technique extremely versatile for a range of applications. Importantly, the advent of PCR created a technical platform for which more advanced and informative methods could be developed. Shown here is the Eppendorf Mastercycler Nexus PCR instrument.
Quantitation
 Although ground-breaking, PCR was originally based upon the concept of the end-point detection. In this non-quantitative format, PCR amplification commences until limiting amounts of reagents are used up in the reaction, at which point the results are measured for the absence or presence of target DNA.
Although ground-breaking, PCR was originally based upon the concept of the end-point detection. In this non-quantitative format, PCR amplification commences until limiting amounts of reagents are used up in the reaction, at which point the results are measured for the absence or presence of target DNA.
Advances in labeling and detection technology meant that true quantitative methods could now be implemented. Monitoring the PCR reaction during the extension or polymerization stage, as opposed to the reaction endpoint, meant that quantitative information could be collected in real-time. Software permitted calculation of actual quantities of DNA in each sample.
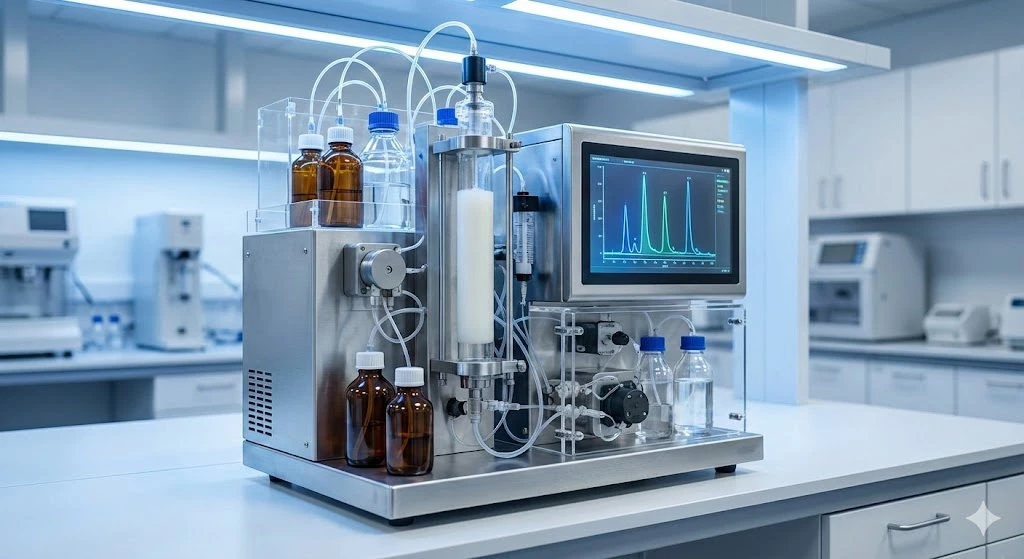
Inclusion of reverse transcriptase upstream of qPCR meant that mRNA levels could now be measured within biological samples, cells, and tissues. The growth of quantitative PCR as a technology enabled inroads for many areas of biomedical research, from gene expression in single cells to rapidly scanning tissues for genetic markers in Immunology and Cancer applications. Shown here is the Bio-Rad CFX96 Touch RT-PCR system.
Scalability
Over time, instrument technologies evolved rapidly on par with the widespread implementation of PCR across disciplines and fields. Instruments and reagents emerged with the ability to detect and quantitate multiple DNA targets in a single sample. Multiple samples could be run on PCR machines with software capable of collecting and deciphering the extensive data. Inevitably, these developments set the stage for the next development – a massive increase in scalability.
Two main factors have enabled the shift in PCR scalability from single reaction to multiplexed digitized performance. The advent of microfluidic technologies meant that samples could now be fractionated into multiple parallel reactions. Implementation of digital technologies enabled readouts that are both highly sensitive and definitive for reactions of this scale.
 Digital PCR (dPCR) emerged as a technique in which the PCR sample is fractionated into individual, parallel reactions – some containing the target DNA and some not. Following PCR cycling reactions, the fraction of those absent target DNA (negative reactions) is used to determine the actual number of target molecules in the sample via Poisson distribution calculations. Importantly, this is without the need for standards or internal controls.
Digital PCR (dPCR) emerged as a technique in which the PCR sample is fractionated into individual, parallel reactions – some containing the target DNA and some not. Following PCR cycling reactions, the fraction of those absent target DNA (negative reactions) is used to determine the actual number of target molecules in the sample via Poisson distribution calculations. Importantly, this is without the need for standards or internal controls.
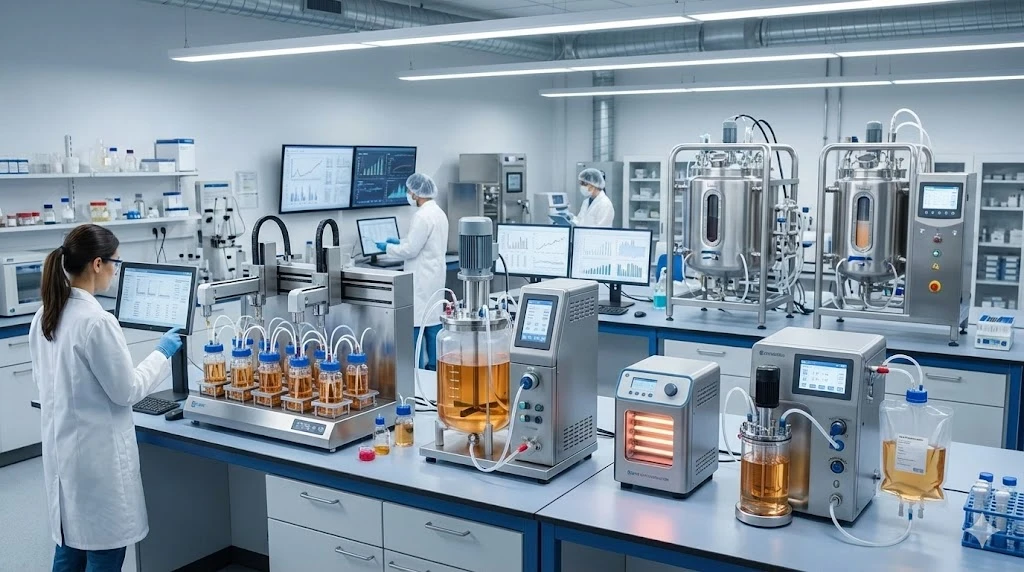
Droplet digital PCR (ddPCR) followed, incorporating microfluidic technology to produce a massive array of thousands of tiny droplets, a few nanoliters in volume, on which digital PCR is then performed. Advantages include digitized detection which avoids the need for labelling or the introduction of additional chemicals and potential detection biases. Microfluidics ensure precision while enabling highly sensitive detection, down to the single cell and single DNA molecule level. The scalability advantage means that procedures which once took hours or days could be performed in single runs of 30 min or less. Shown here is the Bio-Rad QX200 AutoDG Droplet Digital PCR system with automated sample prep component.
Adaptability
Most recently, digital and microfluidic advancements have given rise to chip-based systems designed using semiconductor principles. Unique compact technologies have been developed for lab and remote applications. The technique of isothermal amplification has emerged as well, foregoing the need for the cooling step in thermocycling. A popular version of the technique, looped-mediated isothermal amplification (LAMP), uses a simpler device with lower power consumption than other PCR techniques, and has become a front-runner for remote testing applications.
This brings up a fourth feature of PCR which makes the concept both versatile and powerful – adaptability. Look no further than the recent SARS-CoV-2 outbreak and ensuing pandemic to witness the power of PCR as a platform for critical applications including viral diagnostics and testing.
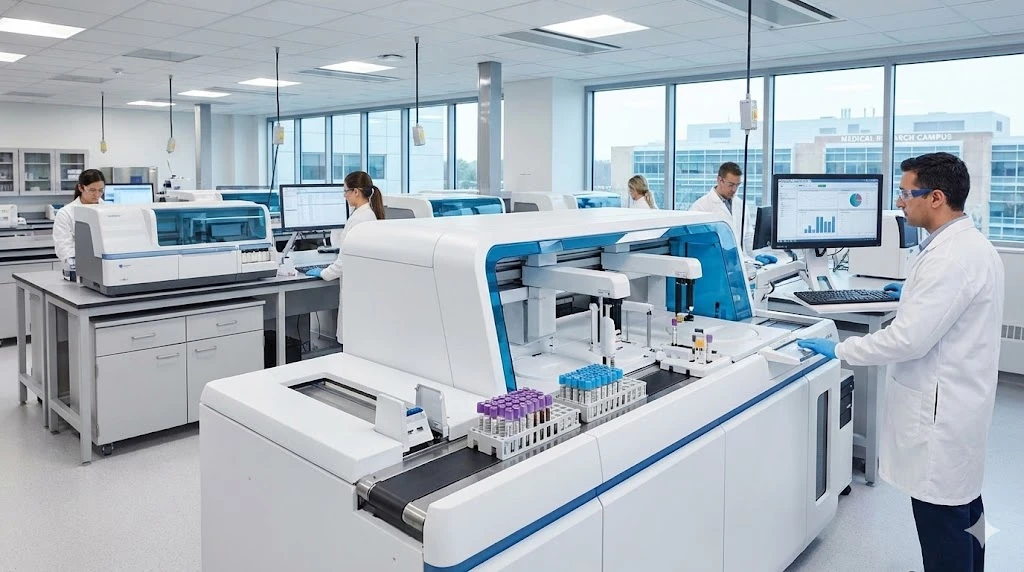
To date there are over 300 FDA-authorized RT-PCR-based diagnostics for SARS-Cov-2 testing. Many are multi-analyte tests which detect more than a single genetic marker, thereby boosting selectivity against false positives and variants that may arise. Others are designed to be compatible with direct-to-consumer sample collection devices or automated sample preparation instruments. Others yet are based around compact, miniaturized, or simplified platforms that offer early detection methods for population-wide monitoring.
The sheer adaptability of PCR has made it invaluable beyond research and clinical diagnostics, into the realm of pandemic preparedness and other critical applications. As instrument technology evolves, it stands to say that PCR will continue to impact our lives far into the future in the lab and beyond.
View PCR, RT-PCR, and Digital PCR Listings
Updated Mar 2023