Ultra-High Resolution Imaging: The New Benchmark in Molecular Microscopy
The world of microscopy has a rich and vivid history, and two recent breakthroughs shine even greater light on an already brilliant field of view.
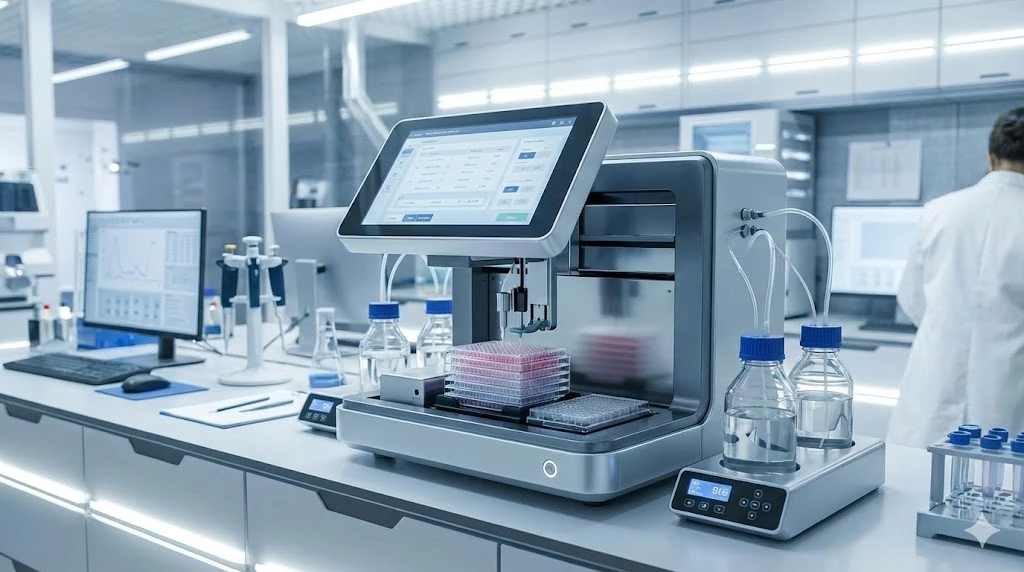
From humble beginnings of the light microscope to modern confocal and electron imaging platforms, the field has progressed by leaps and bounds with developments in technology.
It is now known that the contrast and resolution of optical microscopy imaging can be improved substantially by the use of image deconvolution and computational techniques. Multiple individual images of a sample can be merged computationally to form a superimposed view, breaking the previous limits of resolution for optical technologies. This extra computational power, however, comes with an expense in terms of dataset size and computational resource usage.
A Breakthrough in Image Processing
Recently published in Nature Biotechnology, researchers have now dissected those previous problems into manageable sized bytes with the help of advanced algorithms and software innovations.
- In this work, the authors first present an “unmatched back projector” concept that accelerates deconvolution, relative to commonly used classical algorithms, resulting in a 10-fold increase in image processing.
- They next describe a three-dimensional image-based registration with graphics processing unit which enhances processing speed another 10-100-fold over typical CPU processing.
- Finally, the authors demonstrate that deep learning techniques can provide further acceleration, particularly for deconvolution with spatially varying point spread functions.
What does all this mean? Advancements in microscope imaging have proceeded with substantial gains in resolution and power at the image capture level. The rate-limiting step to faster acquisition has become image deconvolution and post-processing that must occur at the computational level. In other words, the capture of more data than the time to process it – on the scale of terabytes of image data and weeks to months of CPU time.
A big problem is the capture of tiny image blurs, which when compounded can result in a loss of image resolution. The technique of “deconvolution” serves to reduce these blurs, a process that consumes CPU. Compound these CPU costs and, well, the picture becomes clearer.
Stitching many images together to form a superior image in a technique called “parallelization” is another costly process. This was addressed in the current work by breaking the task down and running smaller computations in parallel – then "stitching" the computational results together.
Finally, artificial intelligence (AI) and the concept of neural networking was employed in the workflow in order to focus on the most relevant details, letting the computation work smarter not necessarily harder. In the broader scope, the advancements as a whole move cutting-edge high-resolution fluorescence microscopy within reach of many more scientists across increasing diverse fields of study.
A Breakthrough in Image Resolution
Along the frontier of super-resolution electron microscopy, a recent report in Science Advances has demonstrated improvements in ultra-precise single-molecule microscopy -- heralding a jump forward in molecular imaging power.
The concept of single-molecule localization microscopy (SMLM) wields the potential to map individual molecules and their interactions within intact cells. The techniques, however, are limited by the need for ultra-high precision in localization of the imaging probes and the time span needed for data acquisition. The authors in this work advance the reach of SMLM to now enable distance measurements between molecules on the order of 1-20 nm.
How do they do this? By taking the opposite approach from above -- correcting the image capture precision and obtaining superior data upstream of post-processing steps.
- The researchers first developed an engineering solution called Feedback SMLM, which can capture molecular emissions in a complex and unknown cellular space with equal probability and high-precision. Feedback SMLM performs real-time drift correction in three dimensions, minimizing image signal variation while maxiizing precision.
- Next, they used Feedback SMLM in conjunction with a technique called DNA-PAINT (DNA point accumulation for imaging in nanoscale topography) to accurately measure distances between molecular entities, thus optimizing accuracy.
Together, the approaches circumvented the requirement for elaborate and complex post-processing computational efforts. The result is a stabilized microscope that achieves a stabilization of <1 nm and localization precision of ~1 nm.
What does all of this mean? Building off the 2014 Nobel Prize and the proclamation of the new era of super-resolution microscopy, these new developments now extend our viewing power to the atomic level – several times that of previous limits. Single-molecule investigations just got a lot sharper and potentially much more powerful.
Timely Advancements in Molecular Microscopy
These developments come at a time where the field stands to empower numerous lines of important investigation. The single-molecule resolution advancements described above were tested, proof in concept, on distance measurements of active and resting T cells in situ. The performance of T cell activation in response to infection (called T cell fate) hinges on the quality and quantity of antigen binding to the T cell receptor. Discrete interactions, and the signalling processes they trigger, are essential for a strong and sustained immune response to infection.
Understanding SARS-Cov-2 infection and the development of protective immunity are critical measures in combatting the COVID-19 pandemic. Developments in ultra-high resolution imaging empower the concept of molecular microscopy for the study of viral pathogenesis, cancer, and other important areas -- timely advancements to say the very least.
View microscope and microscope accessories listings on LabX.com